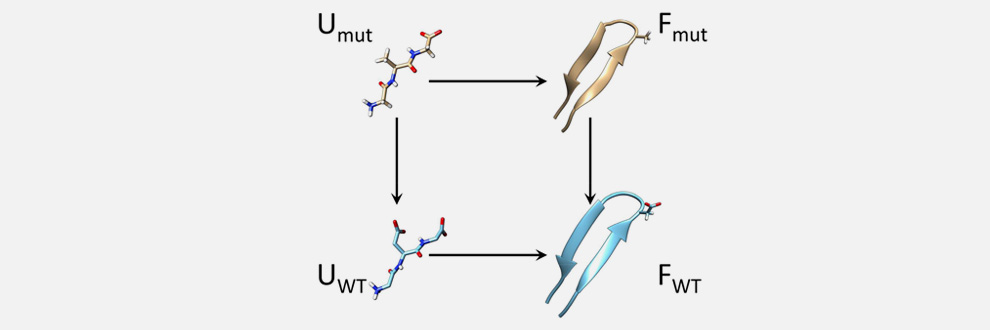
Simulation of Mutational Effects on Peptide Stability
Evaluating the effect of mutations in biomolecules is of paramount importance because these are often associated with diseases. CC&B researchers lead by Guido Tiana, in collaboration with CNR and the University of South Florida, have shown that a recently-developed computational technique can predict the free-energy change upon mutations in a particularly efficient way. The method works when both the wild-type and the mutated molecule display at equilibrium low entropy, that is populate a well-defined structure. The basic idea is to build a thermodynamic cycle involving the two molecules and two “unphysical states” made of unbound atoms constrained by a harmonic potential. The free-energy difference between these two can be calculated analytically, while those between each of the two molecules and the corresponding constrained, unbound atoms can be calculated efficiently by free-energy perturbation. The paper has just been published in J. Phys. Chem. Lett.
Assessment of Mutational Effects on Peptide Stability through Confinement Simulations
Riccardo Capelli, Francois Villemot, Elisabetta Moroni, Guido Tiana, Arjan van der Vaart, and Giorgio Colombo
The Journal of Physical Chemistry Letters Just Accepted Manuscript
DOI: 10.1021/acs.jpclett.5b02221
Available on: pubs.acs.org
